
Introducing Chroma
is a generative model that creates new protein molecules based on geometric and functional programming instructions.
Chroma learns patterns in the three-dimensional structures and amino acid sequences of proteins and protein complexes from the Protein Data Bank. By learning these patterns in a way that generalizes across natural proteins, Chroma can synthesize new protein molecules that adhere to these principles while combining them in novel ways. Importantly, Chroma can be conditioned on a set of desired structural or functional properties, such as the presence of functional structural motifs, symmetry constraints, adhering to a pre-specified shape, belonging to a domain or functional class, or even satisfying text-based descriptions. We think that systems such as Chroma will enable a new, programmable mode of protein engineering where it is routine and feasible to generate specific and tailored protein solutions to complex challenges for bioengineering and human health.
The Chroma system is built on several new machine learning components, including a new neural network architecture for processing and manipulating 3D molecular information, a new diffusion process for adding noise to structures while adhering to the biophysical constraints of protein chains, and a new generalized method for generating high-quality samples from diffusion models. As a result of these innovations, Chroma is able to generate extremely large proteins and protein complexes (e.g. 30,000+ heavy atoms across 4,000+ residues) in a few minutes on a single commodity GPU.

What is a protein?
Proteins are the “doers” of cells, responsible for much of the work that happens in the biological world.
Proteins are formed from a linked sequence of building blocks known as amino acids. Because there are a total of 20 unique “letters” in the amino-acid alphabet, an unimaginably large number of unique proteins are possible, each encoded by a specific sequence of amino acids. While it’s easy to think of proteins as a string of letters, in the cell they fold into 3‑dimensional shapes that perform specific biological functions. A partial list of the feats performed by proteins include replicating DNA, fighting off invading viruses, and keeping your cells alive and healthy. Even things you might not think of as being protein functions, like our sense of smell or sight, are, in fact, enabled by proteins! Given how capable this molecular class is, it is not surprising that most of the medicines approved today are, in fact, proteins. Thus, the better we understand how proteins work and the more effectively that we can generate new ones for targeted functions, the more effective will be the medicines of tomorrow.
Sampling from the protein universe
Having learned common principles from natural proteins, Chroma can generate random proteins without any additional prompting.
Below is a small set of single-chain proteins sampled from the model.

Sampling protein complexes
Many proteins carry out their functions through interactions with other proteins in multi-molecule assemblies called complexes.
These complexes can transmit information, catalyze important chemical reactions, or act as complex molecular machines and are frequently the targets of therapies. Chroma can directly generate protein complexes composed of many proteins of different shapes. Below are a few examples of complexes generated by Chroma.

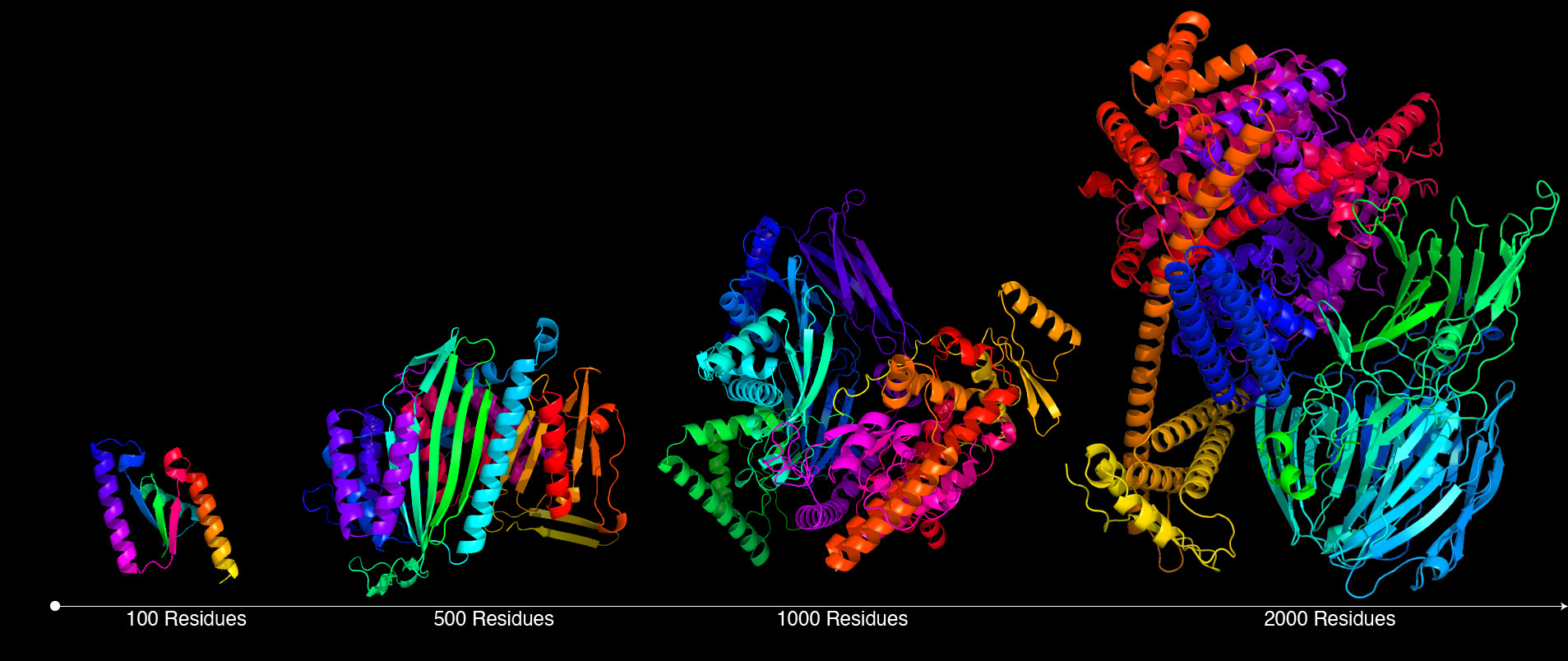
Making giants
Typical proteins are composed of tens to thousands of amino acids. Chroma’s efficient computational scaling makes it possible to directly generate large molecular assemblies at the scales frequently seen in nature.
Programming proteins with Chroma
Chroma has the ability to incorporate a wide range of properties and constraints to steer the generative process. We imagine that capabilities like this will enable a future in which scientists can specify desired protein functions or properties in a high-level language and allow Chroma to compile these attributes in a lower-level “executable” version of the molecular function in the form of a 3D protein molecule.
In software engineering, the transition from low-level machine and assembly codes to high-level languages such as C++/Java/Python sparked an intense period of innovation in which developers could build complicated programs from simpler and reliable abstractions. In a similar way, we believe that generative models such as Chroma offer a step towards a higher level programming abstraction for biology. Below we show an early version of what we imagine this could look like.
Symmetry groups
Many protein complexes in nature are built from symmetrical “tilings” of one or more protein building blocks.
Chroma can be conditioned on many different kinds of symmetry, from simple circular symmetries to the complex icosahedral symmetries often seen in nanoparticles.
Protein infilling
A routine challenge in protein engineering is to keep part of a protein molecule that is important for one property (such as folding and interacting with the immune system) fixed while changing another part of the protein (e.g. to bind to a target).
In the example below an antibody heavy chain CDR region is being reimagined by the model. The top left image shows a true structure of an antibody-protein complex. In the remaining 3 images, the loops are removed and re-generated from scratch.

Semantic conditioning
Chroma can be conditioned by other neural networks that know about proteins without retraining. This makes it possible to control higher-level properties of properties, i.e. their semantics.
In the example below, we bias Chroma sampling with two different neural networks, one which was trained to predict CATH folds from structure, and the other of which was trained to predict natural language captions from protein 3D structures. Each column represents a particular conditioning example. The leftmost two columns are conditioned with the CATH classification model. The rightmost two columns are conditioned with the protein captioning model. The top row of structures are random samples that the model generated without conditioning. The middle row is the sample drawn from Chroma with conditioning. Finally, the bottom row shows a real example belonging to the desired class or caption. We can see how the classifier drives the samples (middle row) to look more similar to real examples (bottom row) than unconditioned (top row). Capabilities such as this should make high level functional programming more routinely feasible.

Shape control
What are the limits on protein shape? We can also ask Chroma to sample 3D structures given arbitrary shape specifications. Below we asked for proteins consistent with the Latin alphabet and numeral system.
Transforming between protein structures
Since Chroma parameterizes the space of possible 3D protein structures in a continuous way, we can ask what it thinks is in between two different structures.
These morphs shed light on how Chroma organizes structure space in its internal representations.

Morphing between secondary structures
We morph between a highly alpha-helical protein (pheromone from marine ciliate Euplotes raikovi, PDB 6E6N) and a beta-rich protein (toxin from scorpion Mesobuthus martensii PDB 6AY8).

A bigger alpha to beta morph
This morph transforms between rhodopsin, a transmembrane protein found in rod cells in the eye (PDB 2I35), and a six-bladed beta propeller structure (PDB 3DAS).

Parallel-to-anti-parallel transition
Coiled coils are a common structural motif involving alpha-helices that wind around one another. This morph shows a transition between a parallel (PDB 2ZTA) and an anti-parallel coiled coil (PDB 1HF9).

Conformational shifts in a transporter
Leucine transporter (LeuT) helps move the amino acid leucine across cell membranes via a dynamics conformational change of opening and closing. Here we interpolate between three experimentally-determined conformations: the outward-open state (PDB 3TT1), the occluded state (PDB 3F3E), and the inward-open state (PDB 3TT3).

Viral fusion-enabling conformational shifts
Hemagglutinins are spike-shaped proteins on the surfaces of viruses such as influenza that drive fusion of the virus with the host cell. Here we morph between three different functional conformational states of this fusion process of Influenza hemagglutinin: state I (PDB 6Y5H), state II (PDB 6Y5I), and state IV (PDB 6Y5K). We can see the interior alpha helices retract to drive fusion with the host membrane.
The Details

This model was made possible by several novel components
- Programable protein generation via a collection of new conditioning models
- Random Graph Neural Networks. A novel neural network architecture that can process and modify 3D molecular systems in time that scales sub-quadratically with the whole system.
- Polymer diffusion. A novel diffusion process that respects the biophysical constraints of proteins as collapsed polymers
- Low-temperature Sampling. A novel sampling algorithm for generating high-likelihood samples from diffusion models

Structural validation
Across a set of 10,000 samples of single-chain structures, we find that Chroma reproduces the structural statistics of proteins from the PDB.
Designability
Chroma generates protein molecules by first synthesizing a backbone structure and then designing sequences consistent with those 3D backbones. While the only true way to test the validity of these designs is experimental characterization, we can begin to check self-consistency by asking an orthogonal in silico structure prediction method whether it thinks the designed sequence should fold back into its intended shapes.
We find frequent agreement, even for reasonably sized molecules shown below. Of course, the true test of protein design is to make and test molecules in the lab, which is why machine learning is only the first step of what we do at Generate.

Novelty
Natural proteins tend to be composed of well-defined conserved structural domains ranging between 50 and 200 residues in length. We assess novelty of Chroma-generated structures by computing the required number of common protein domains (CATH) needed to cover at least 80% of each generated structures above a structural cutoff (TM > 0.5). By this measure, Chroma proteins would seem to demonstrate greater novelty than collection of proteins from the PDB, despite being trained on proteins from the PDB. This could be a sign of learning the principles required for generalization, while not overfitting or memorizing the proteins that have already been seen.
Limitations

Examples of generated proteins that did not work well
Above are some examples of generated proteins that may illustrate some potential bugs and failure modes. (a) In some cases the conditioners result in protein samples that are not connected or are very sparsely connected. (b) Unconditioned samples can exhibit rare but significant pathologies such as clashes, poor topologies, and tangles. Large unconditioned samples (1000+ residues) sometimes have extended regions with low secondary structure content.
Some limitations are:
- The real test of any protein is wet lab synthesis and experimentation, and so far these are only in silico predictions.
- Conditional models can be difficult to tune and still tend to require expert supervision and collaboration with experienced protein designers for troubleshooting.
- Low temperature sampling can adjust macroscopic observables such as the balance of alpha and beta secondary structure content, which requires deeper understanding.
Read the paper
Illuminating protein space with a programmable generative model
John Ingraham, Max Baranov, Zak Costello, Vincent Frappier,
Ahmed Ismail, Shan Tie, Wujie Wang, Vincent Xue, Fritz Obermeyer, Andrew Beam, Gevorg Grigoryan
Three billion years of evolution have produced a tremendous diversity of protein molecules, and yet the full potential of this molecular class is likely far greater. Accessing this potential has been challenging for computation and experiment because the space of possible protein molecules is much larger than the space that are likely to host function. Here we introduce Chroma, a generative model for proteins and protein complexes that can directly sample novel protein structures and sequences and that can be conditioned to steer the generative process towards desired properties and functions. To enable this, we introduce a diffusion process that respects the conformational statistics of polymer ensembles, an efficient neural architecture for molecular systems based on random graph neural networks that enables long-range reasoning with sub-quadratic scaling, equivariant layers for efficiently synthesizing 3D structures of proteins from predicted inter-residue geometries, and a general low-temperature sampling algorithm for diffusion models. We suggest that Chroma can effectively realize protein design as Bayesian inference under external constraints, which can involve symmetries, substructure, shape, semantics, and even natural language prompts. With this unified approach, we hope to accelerate the prospect of programming protein matter for human health, materials science, and synthetic biology.
We would like to thank William F. DeGrado and Generate employees Adam Root, Alan Leung, Alex Ramos, Brett Hannigan, Eugene Palovcak, Frank Poelwijk, James Lucas, James McFarland, Karl Barber, Kristen Hopson, Martin Jankowiak, Mike Nally, Molly Gibson, Ross Federman, Stephen DeCamp, Thomas Linsky, Yue Liu, and Zander Harteveld for reading of the manuscript draft and providing helpful comments.